Software
Computational Tools
RNA Analysis Software
Open-source tools for RNA variant discovery, RNA editing, and transcriptome-scale analysis.
I develop production-ready bioinformatics methods in Python, including packaging and distribution workflows for reproducible research software. I am also proficient in R and Bash for scalable analysis pipelines and rapid prototyping.
View GitHub ProfileisoLASER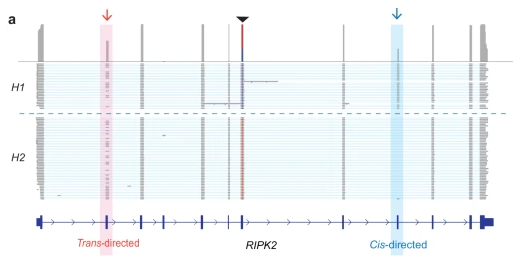
Role: Lead developer
Isoform-aware analysis workflows for long-read transcriptomics and RNA processing.
scAllele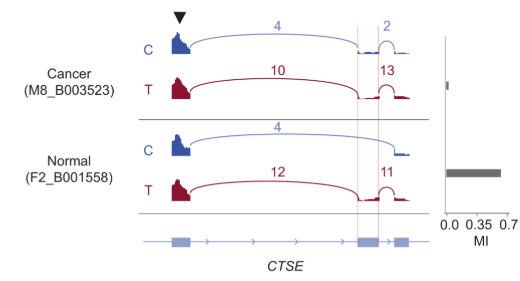
Role: Lead developer
Variant detection and downstream analysis from single-cell RNA sequencing data.
L-GIREMI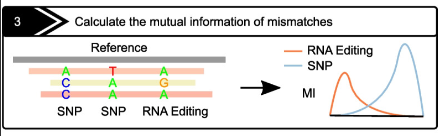
Role: Contributor
Long-read RNA editing site identification for transcriptome-wide studies.
BEAPR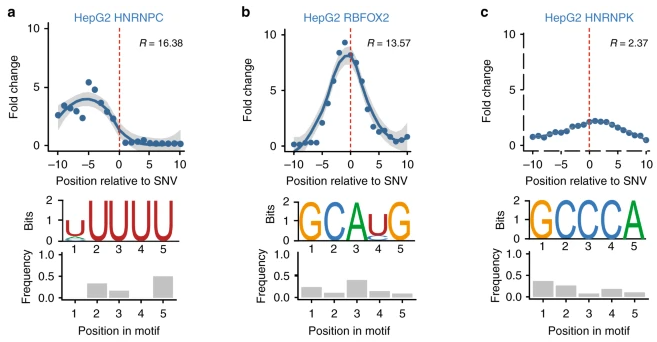
Role: Contributor
Computational framework for identifying functional allele-specific molecular signals.

